Welcome
 |
Radhey S. Gupta (Professor)
Department of Biochemistry and Biomedical Sciences (HSC- 4H2)
1200 Main Street West,
Hamilton, Ontario, Canada L8N 3Z5
Phone: (905) 525-9140 ex. 22639
Fax: (905) 522-9033
Email: gupta@mcmaster.ca |
Comparative Genomics:
Microbial Evolution, Diagnostics and Functional Studies
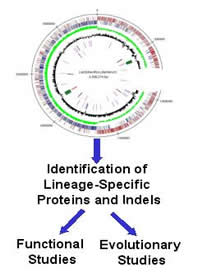 Rare Genetic Changes (RGCs), which are uniquely shared by species from well-defined groups of organisms, provide powerful molecular markers of common evolutionary descent for evolutionary, functional and diagnostic studies. The main focus of our current research work is to use sequence information from different genomes to identify unique molecular characteristics, such as signature proteins or conserved indels (i.e. inserts or deletions) in protein sequences, that are distinctive characteristics of different groups of organisms at various taxonomic levels (viz. species and strains, genus, order, family or whole phylum). These novel molecular characteristics provide us with powerful tools for identifying different groups of organisms in clear terms and for a variety of investigations. Using these rare genomic changes, work is being carried out in a number of different areas. These include (i) understanding the evolutionary relationships among different prokaryotic and eukaryotic organisms; (ii) Understanding the cellular functions of these lineage-specific signature proteins as well as lineage-specific conserved inserts and deletions in important housekeeping proteins by genetic and biochemical studies; (iii) Development of novel diagnostic methods (PCR based and immunological) for identification of different groups of organisms based upon these signature proteins and conserved indels; (iv) The use of these lineage-specific probes with predicitive ability to identify/explore the presence of different groups of organisms in metagenomic sequences from various environments. Although our work in this area has thus far mainly focused on prokaryotic organisms, this approach is now being extended to the study of eukaryotic organisms. For additional information and recent publications from my lab in this area see Publications and the Bacterial Phylogeny website. Rare Genetic Changes (RGCs), which are uniquely shared by species from well-defined groups of organisms, provide powerful molecular markers of common evolutionary descent for evolutionary, functional and diagnostic studies. The main focus of our current research work is to use sequence information from different genomes to identify unique molecular characteristics, such as signature proteins or conserved indels (i.e. inserts or deletions) in protein sequences, that are distinctive characteristics of different groups of organisms at various taxonomic levels (viz. species and strains, genus, order, family or whole phylum). These novel molecular characteristics provide us with powerful tools for identifying different groups of organisms in clear terms and for a variety of investigations. Using these rare genomic changes, work is being carried out in a number of different areas. These include (i) understanding the evolutionary relationships among different prokaryotic and eukaryotic organisms; (ii) Understanding the cellular functions of these lineage-specific signature proteins as well as lineage-specific conserved inserts and deletions in important housekeeping proteins by genetic and biochemical studies; (iii) Development of novel diagnostic methods (PCR based and immunological) for identification of different groups of organisms based upon these signature proteins and conserved indels; (iv) The use of these lineage-specific probes with predicitive ability to identify/explore the presence of different groups of organisms in metagenomic sequences from various environments. Although our work in this area has thus far mainly focused on prokaryotic organisms, this approach is now being extended to the study of eukaryotic organisms. For additional information and recent publications from my lab in this area see Publications and the Bacterial Phylogeny website.
Previously, my lab has also carried out extensive work on i) mitochondrial and heat shock proteins and their presence and functions at extra-mitochondrial locations, and ii) on the structure and function of mammalian Adenosine Kinase . However, these areas are currently not actively worked on in my lab. Please refer to publications for our work in these areas. |

